1. What types of molecules can be quantified by Two-Tailed qPCR?
Our patented TT-qPCR platform has superior performance in the detection and quantification of short sequence RNA molecules (<300bp). This includes short and mid-sized non-coding RNAs (sncRNAs & mncRNAs) like microRNAs, Piwi-interacting RNAs, snoRNAs and others. In various diagnostic panels, it is possible to combine more types of targets, e.g., piRNAs and miRNAs.
2. How many samples does one Two-Tailed PRIMER kit measure?
The basic TT-PRIMER kit includes reverse transcription (RT) primer for 50 samples and two qPCR primers (forward and reverse) for 150 reactions as we believe in golden standard of running PCR in a triplicate per sample.
3. Which products shall I purchase to run this assay?
Complete TT-qPCR assay includes Two-Tailed PRIMERs (RDTT000****PRI), Two-Tailed cDNA Synthesis System 50rxn (RDTTRT50) and Two-Tailed qPCR Master Mix 150rxn (RDTTPCR150). There is also a possibility to order a synthetic miRNA (TTK0000****) with various applications, e.g., isolation spike-in control, positive control, absolute quantification etc.
4. Can I multiplex several targets in one reaction?
Yes, there is a possibility to multiplex up to 5 targets within one RT reaction. This approach is very sequence dependent, and our specialists perform laboratory tests to verify there is no cross reactivity or alterations in assays characteristics like sensitivity or specificity. Following PCR reaction always runs separately for each target as the detection is through SYBR Green dye.
5. What laboratory equipment is necessary to run the Two-Tailed qPCR?
Our assays can be run in any laboratory equipped for standard qPCR measurement. You need a vortex mixer and benchtop centrifuge, standard block heater or thermocycler and Real-Time PCR system with FAM/SYBR Green detection channel.
6. Could I order primers for ANY microRNA targets?
YES, besides already developed assays (list of them at www.biovendor.com/two-tailed-rt-qpcr) we offer also customized development and production of TT-PRIMERs. So whatever miRNA, piRNA, siRNA, SNORD or other sncRNA/mncRNA you are interested in, we will develop a unique, specific, and highly sensitive assay to quantify it. Development time is approximately 6-8 weeks. It can be altered in case of additional testing, e.g., for multiplex reaction.
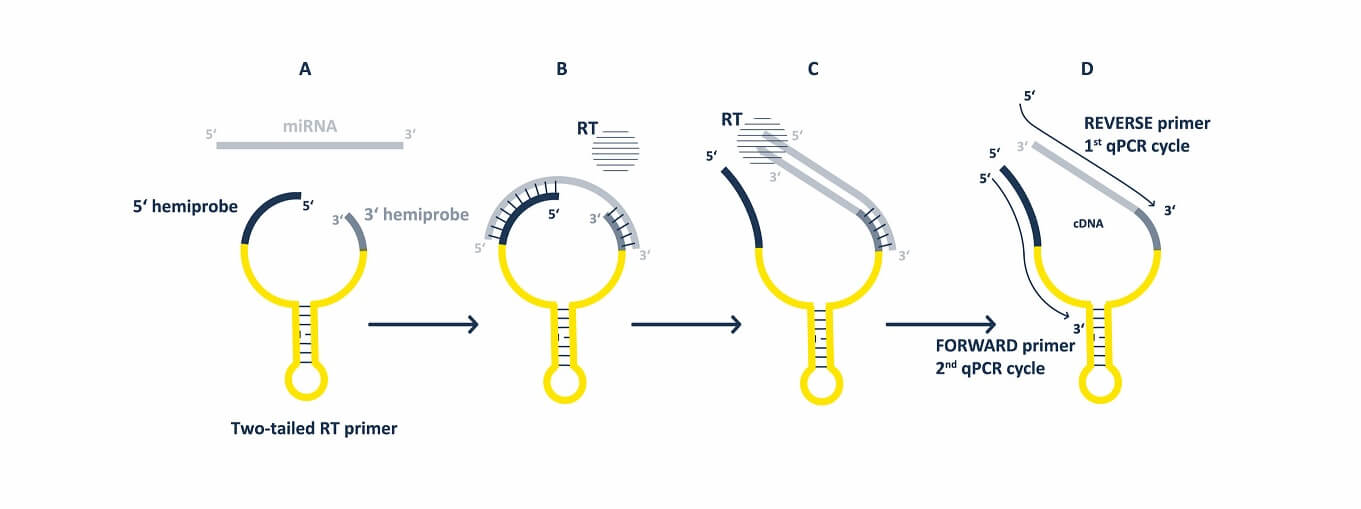
7. How is a TT-qPCR assay developed?
Our team of specialists always starts with in-silico design of all primers using a comprehensive algorithm. Our unique Two-Tailed RT primers are then laboratory tested together in several combinations with various qPCR primers to identify the optimal combination of RT and qPCR primers which meets all our requirements for an assay performance. That means that we always run various laboratory tests to make sure that the assays quantify only desired targets.
8. What type of controls and normalisation shall I use for my project?
There are several options how to properly approach normalisation and what controls to include in your research. Regarding controls, besides including the blank negative control of RT reaction and PCR reaction, we suggest usage of a positive control (synthetic miRNA molecule – available at www.biovendor.com/two-tailed-rt-qpcr). It is always wise to control very important step, the isolation of RNA. Additionally, we prepared an isolation control kit which contains an artificial miRNA in the combination with a specific C.elegans miRNA. These two spike-ins are used to monitor the isolation efficiency. This can be further used also as an exogenous normalization and mostly it works as a comparison between each sample which enters the TT-qPCR protocol. You can also choose a stable expressed sncRNA target known to be present in your particular sample type which works as an endogenous normalization control. Last option is to prepare a dilution set of a synthetic miRNA of your choice and measure it as a calibration curve for an absolute quantification of your target.
9. What isolation kits shall I use to obtain ideal performance?
There are many producers available on the market and it is always dependent on your experiences but, our assays are tested and validated with RNA isolates eluted from our own isolation kits (www.biovendor.com/mirna-isolation-kits). They are all column-based, sample type specific, user friendly with very attractive and competitive price. Our specialists are always willing to provide a consultation, or any troubleshooting based on your individual project settings. When you are planning your project, it is very important to use the same isolation procedure across all your experiments you would like to compare. Be aware of using and combining samples proceeded by different isolation kits.
10. What sample types can be measured by TT-qPCR assay?
TT-qPCR assay can measure RNA isolated from any type of biological material (tissues, cell lines, solid tumours, etc.) and biofluids (blood, serum, plasma, urine, sweat, etc.). The detection of the specific target in your biological sample depends on its concentration in the sample, so we recommend testing the capture in the small preliminary study. In the case of customised assay development, our specialist can test the capture in your specific samples during the development of the assay.
